Ensemble learning
Using a single model can not achieve the best results, so we consider using ensemble learning method to further reduce the error.
Integrated learning is the stacking and integration of different models and the selection of optimal parameters.
Thirteen models will be used in ensemble learning. First, import the packages that need to be used.
from sklearn.model_selection import cross_val_score, GridSearchCV, KFold from sklearn.linear_model import LinearRegression from sklearn.linear_model import Ridge from sklearn.linear_model import Lasso from sklearn.ensemble import RandomForestRegressor, GradientBoostingRegressor, ExtraTreesRegressor from sklearn.svm import SVR, LinearSVR from sklearn.linear_model import ElasticNet, SGDRegressor, BayesianRidge from sklearn.kernel_ridge import KernelRidge from xgboost import XGBRegressor
1. Basic Modeling and Evaluation
According to the requirements of the competition, the cross validation evaluation index based on RMSE is defined at first.
#Define cross-validation strategies and evaluation functions def rmse_cv(model,X,y): rmse = np.sqrt(-cross_val_score(model, X, y, scoring="neg_mean_squared_error", cv=5)) return rmse
Save all the models to be used
models = [LinearRegression(),Ridge(),Lasso(alpha=0.01,max_iter=10000),RandomForestRegressor(),GradientBoostingRegressor(),SVR(),LinearSVR(), ElasticNet(alpha=0.001,max_iter=10000),SGDRegressor(max_iter=1000,tol=1e-3),BayesianRidge(),KernelRidge(alpha=0.6, kernel='polynomial', degree=2, coef0=2.5), ExtraTreesRegressor(),XGBRegressor()]
Firstly, 13 models and 5-fold cross-validation are used to evaluate the prediction effect of each model.
names = ["LR", "Ridge", "Lasso", "RF", "GBR", "SVR", "LinSVR", "Ela","SGD","Bay","Ker","Extra","Xgb"] for name, model in zip(names, models): score = rmse_cv(model, X_scaled, y_log) print("{}: {:.6f}, {:.4f}".format(name,score.mean(),score.std()))
LR: 621771864740.772827, 411989810656.7503 Ridge: 0.118922, 0.0076 Lasso: 0.118914, 0.0065 RF: 0.148033, 0.0049 GBR: 0.123131, 0.0076 SVR: 0.179019, 0.0129 LinSVR: 1.239646, 0.4882 Ela: 0.116366, 0.0070 SGD: 2.814280, 0.5916 Bay: 0.117589, 0.0066 Ker: 0.114100, 0.0081 Extra: 0.141861, 0.0112 Xgb: 0.124880, 0.0058
2. Adjusting the parameters of each model
Establish a parameter adjustment method, where the evaluation index is RMSE, so the printed score should also be RMSE. Define the crossover mode, specify the model first and then specify the parameters, which is convenient to test multiple models, and use grid cross validation.
class grid(): def __init__(self,model): self.model = model def grid_get(self,X,y,param_grid): grid_search = GridSearchCV(self.model,param_grid,cv=5, scoring="neg_mean_squared_error") grid_search.fit(X,y) print(grid_search.best_params_, np.sqrt(-grid_search.best_score_)) grid_search.cv_results_['mean_test_score'] = np.sqrt(-grid_search.cv_results_['mean_test_score']) print(pd.DataFrame(grid_search.cv_results_)[['params','mean_test_score','std_test_score']])
Result of Lasso()
grid(Lasso()).grid_get(X_scaled,y_log,{'alpha': [0.0004,0.0005,0.0007,0.0006,0.0009,0.0008],'max_iter':[10000]})
{'alpha': 0.0009, 'max_iter': 10000} 0.11557402177546283
params mean_test_score std_test_score
0 {'alpha': 0.0004, 'max_iter': 10000} 0.116897 0.001659
1 {'alpha': 0.0005, 'max_iter': 10000} 0.116580 0.001644
2 {'alpha': 0.0007, 'max_iter': 10000} 0.116041 0.001612
3 {'alpha': 0.0006, 'max_iter': 10000} 0.116301 0.001630
4 {'alpha': 0.0009, 'max_iter': 10000} 0.115574 0.001574
5 {'alpha': 0.0008, 'max_iter': 10000} 0.115794 0.001591
Result of Ridge()
grid(Ridge()).grid_get(X_scaled,y_log,{'alpha':[35,40,45,50,55,60,65,70,80,90]})
{'alpha': 90} 0.11753822142197719
params mean_test_score std_test_score
0 {'alpha': 35} 0.118097 0.001621
1 {'alpha': 40} 0.118003 0.001607
2 {'alpha': 45} 0.117921 0.001595
3 {'alpha': 50} 0.117849 0.001583
4 {'alpha': 55} 0.117787 0.001573
5 {'alpha': 60} 0.117733 0.001564
6 {'alpha': 65} 0.117686 0.001555
7 {'alpha': 70} 0.117646 0.001547
8 {'alpha': 80} 0.117582 0.001533
9 {'alpha': 90} 0.117538 0.001522
After many rounds of testing, the following six models and their corresponding optimal parameters are finally selected.
lasso = Lasso(alpha=0.0005,max_iter=10000) ridge = Ridge(alpha=60) svr = SVR(gamma= 0.0004,kernel='rbf',C=13,epsilon=0.009) ker = KernelRidge(alpha=0.2 ,kernel='polynomial',degree=3 , coef0=0.8) ela = ElasticNet(alpha=0.005,l1_ratio=0.08,max_iter=10000) bay = BayesianRidge()
3. Integration method, using weighted averaging
Weighted average of each model according to weight
##Defining a weighted average is equivalent to writing fit_transform() class AverageWeight(BaseEstimator, RegressorMixin): def __init__(self,mod,weight): self.mod = mod##Number of models self.weight = weight##weight def fit(self,X,y): self.models_ = [clone(x) for x in self.mod] for model in self.models_: model.fit(X,y) return self def predict(self,X): w = list() # pred returns the cross-validation results for each model in size (number of models) x (number of samples in validation set) pred = np.array([model.predict(X) for model in self.models_]) # For each data point, a single model is multiplied by weights and then added up. for data in range(pred.shape[1]): single = [pred[model,data]*weight for model,weight in zip(range(pred.shape[0]),self.weight)] w.append(np.sum(single)) return w
Define six initial weights
w1 = 0.02 w2 = 0.2 w3 = 0.25 w4 = 0.3 w5 = 0.03 w6 = 0.2
weight_avg = AverageWeight(mod = [lasso,ridge,svr,ker,ela,bay],weight=[w1,w2,w3,w4,w5,w6]) print(rmse_cv(weight_avg,X_scaled,y_log)) print(rmse_cv(weight_avg,X_scaled,y_log).mean())##Calculate the mean of cross validation
[0.11579461, 0.12567714, 0.12194691, 0.10298115, 0.11307847] 0.11589565350500453
4. stacking model stacking
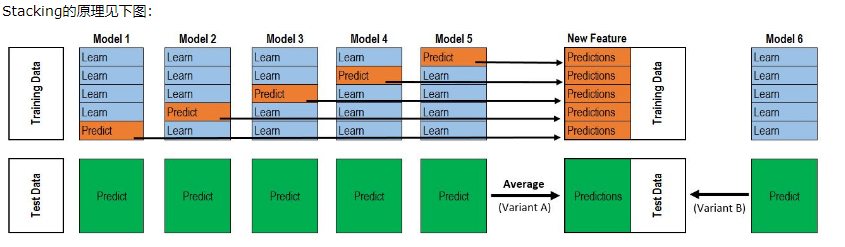
class stacking(BaseEstimator, RegressorMixin, TransformerMixin): def __init__(self,mod,meta_model): self.mod = mod self.meta_model = meta_model#Meta model self.kf = KFold(n_splits=5, random_state=42, shuffle=True)##That's the biggest feature of stacking. It's broken down a few times. def fit(self,X,y): self.saved_model = [list() for i in self.mod] # Preservation model oof_train = np.zeros((X.shape[0], len(self.mod))) # Here we get a matrix of the number of rows multiplied by the number of models in the training set. for i,model in enumerate(self.mod):#What is returned is the index and the model itself. for train_index, val_index in self.kf.split(X,y):##The data returned is from the province. renew_model = clone(model)##Model replication renew_model.fit(X[train_index], y[train_index])#Training data self.saved_model[i].append(renew_model)##Add the model in oof_train[val_index,i] = renew_model.predict(X[val_index])##Used to predict validation sets self.meta_model.fit(oof_train,y)#Meta model return self def predict(self,X): whole_test = np.column_stack([np.column_stack(model.predict(X) for model in single_model).mean(axis=1) for single_model in self.saved_model]) ##What you get is the entire test suite. return self.meta_model.predict(whole_test)#What is returned is to use the metamodel to predict the entire test set. def get_oof(self,X,y,test_X): oof = np.zeros((X.shape[0],len(self.mod)))##Initialization is 0 test_single = np.zeros((test_X.shape[0],5))##Initialization is 0 test_mean = np.zeros((test_X.shape[0],len(self.mod))) for i,model in enumerate(self.mod):##i is a model. for j, (train_index,val_index) in enumerate(self.kf.split(X,y)):##j is all partitioned data clone_model = clone(model)##Cloning module is equivalent to copying the model. clone_model.fit(X[train_index],y[train_index])##Training the segmented data oof[val_index,i] = clone_model.predict(X[val_index])##Predicting Verification Sets test_single[:,j] = clone_model.predict(test_X)##Predicting test sets test_mean[:,i] = test_single.mean(axis=1)##Test Setting Well Means return oof, test_mean
##After preprocessing, it can be placed in a stacked model for calculation. a = Imputer().fit_transform(X_scaled)#Equivalent to x b = Imputer().fit_transform(y_log.values.reshape(-1,1)).ravel()#Equivalent to y
stack_model = stacking(mod=[lasso,ridge,svr,ker,ela,bay],meta_model=ker) stack_model.fit(a,b)#Model training pred = np.exp(stack_model.predict(test_X_scaled))#Forecast result.to_csv("submission2.csv",index=False)
The post-upload score was 0.12310