catalogue
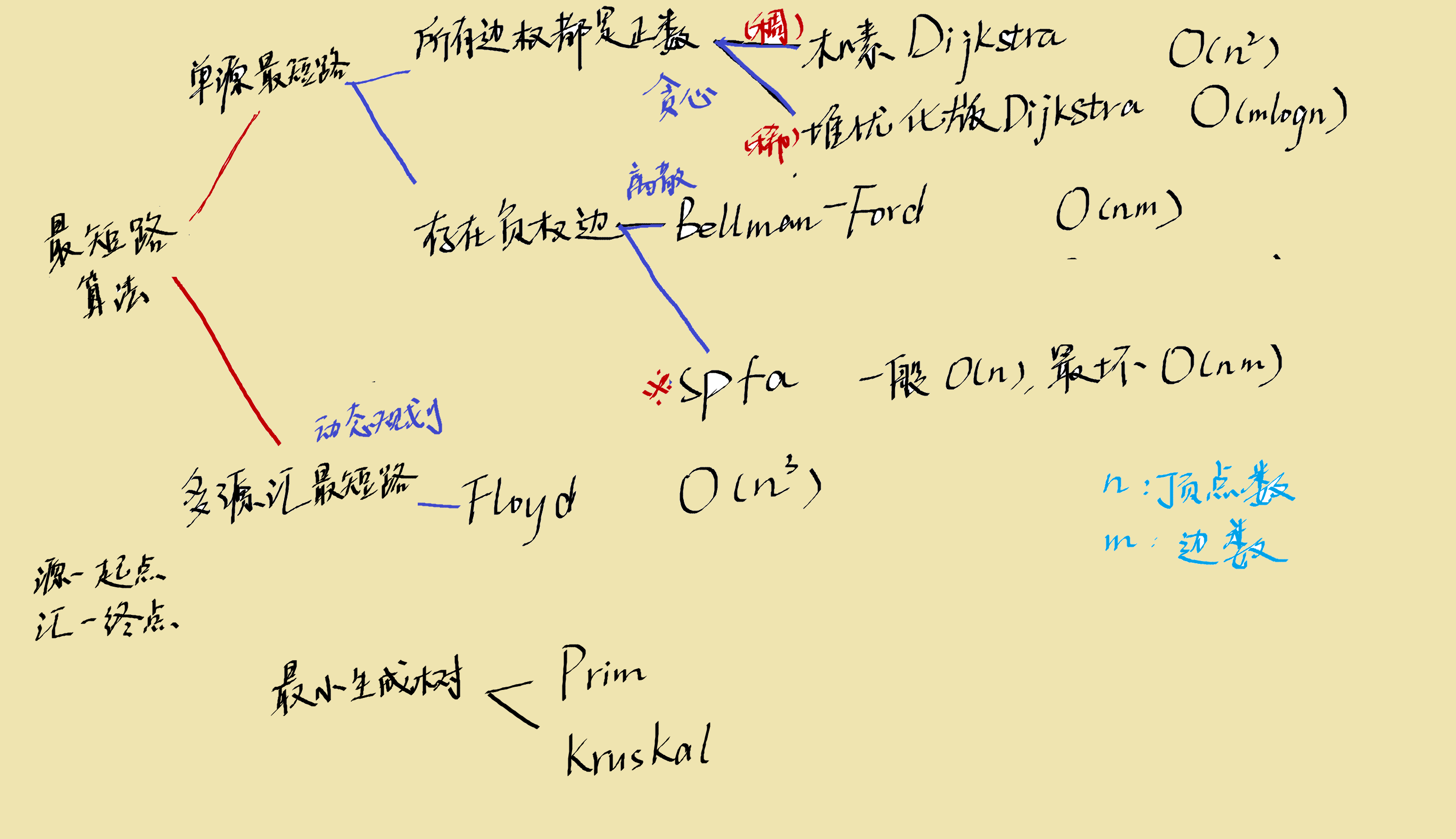
shortest path
Plain Dijkstra
Time complexity: O(n2+m) , n Represents the number of points, M Represents the number of sides
Dense graph 👉 Storage form: adjacency matrix
Template: g[x][y] stores the edge weight during initialization, st[i] indicates whether I is added to the gradually expanding set, dist[i] What we save is the shortest distance from 1 to I, which is our goal
Idea:
Seek 1 → n Shortest path
one Initialization distance dist[1] = 0, dist[i] = +∞
2. for i 1~n:
for j 1~n:
t ← Not in set s, The point closest to the set
s ← t (join the organization, i.e. st[t]=true)
for j 1~n:
Use t to update the distance of other points (the distance from the points already in the set to 1 may not be the shortest, and the dist [J] value is updated to min(dist[j], dist[t] + g[t][j]) )
Input example:
3 3 1 2 2 2 3 1 1 3 4
Output example:
3
#include <iostream>
#include <cstring>
#include <algorithm>
using namespace std;
const int N = 510;
int n, m;
int dist[N],g[N][N];
bool st[N]; //Whether the tag is in the collection
int dijkstra()
{
memset(dist, 0x3f, sizeof dist);
dist[1] = 0;
for(int i = 0; i < n; i ++)
{
int t = -1;
for(int j = 1; j <= n; j ++)
if(!st[j] && (t==-1 || dist[j] < dist[t]))
t = j;
//if(t == n) break; // You can break in advance to optimize the time overhead
st[t] = true;
for(int j = 1; j <= n; j ++) //Update the shortest path in the collection
dist[j] = min(dist[j], dist[t]+g[t][j]);
}
if(dist[n] == 0x3f3f3f3f) return -1;
else return dist[n];
}
int main()
{
cin >> n >> m;
memset(g, 0x3f, sizeof g);
int a,b,c;
for(int i = 0; i < m; i++)
{
scanf("%d%d%d", &a, &b, &c);//More edges, scanf saves time
g[a][b] = min(c, g[a][b]); //According to the meaning of the question, there may be double edges and self rings in the figure
}
cout << dijkstra();
}minimum spanning tree
Prim algorithm
(very similar to Dijkstra's idea, there is only difference when updating dist, the former is the point with the smallest distance from the set, and the latter is the point with the smallest distance from the starting point)
Idea:
one Initialization distance dist[i] = +∞
2. for i 1~n:
for j 1~n:
t ← Not in set s, The point closest to the set
If I= 1 (i.e. not the point at the initial time) and dist[t] = = + ∞ (the graph is not connected), Exit directly (supplement: as long as it is not the first point, dist[t] represents the length between the current point and a point in the connected set)
for j 1~n:
Use t to update the distance of other points (the distance from the points already in the set to 1 may not be the shortest, and the dist [J] value is updated to min(dist[j], g[t][j]) (supplement: min(dist[j], g[t][j]), dist [J] indicates that when point t is not added to the set before Distance from point J to set, g[t][j] Indicates if the point t is added to the set Distance from point J to set)

Input example:
4 5 1 2 1 1 3 2 1 4 3 2 3 2 3 4 4
Output example:
6
#include <iostream>
#include <cstring>
#include <algorithm>
using namespace std;
const int N = 510, INF = 0x3f3f3f3f;
int n,m;
int g[N][N],dist[N];
bool st[N];
int prim()
{
memset(dist, INF, sizeof dist);
int res = 0; //Sum of weights of minimum spanning tree
for(int i = 0; i < n; i++)
{
int t = -1;
for(int j = 1; j <= n; j++)
if(!st[j] && (t == -1 || dist[j] < dist[t]))
t = j;
if(i && dist[t]==INF) return INF;
if(i) res += dist[t];//★ add before updating, otherwise the weight of self ring may be included
for(int j = 1; j <= n; j++)
dist[j] = min(dist[j], g[t][j]);//★
st[t] = true;//★
}
return res;
}
int main()
{
cin >> n >> m;
memset(g, INF, sizeof g);
int a, b, c;
while (m -- )
{
scanf("%d%d%d", &a, &b, &c);
g[a][b] = g[b][a] = min(c, g[a][b]); //★
}
int res = prim();
if(res == INF) cout << "impossible";
else cout << res;
return 0;
}Kruskal algorithm
(in essence, it is a joint query set)
Idea:
one All edges are sorted by weight from small to large
two Enumerate each edge ab, Weight c
If a and B are not connected, add this edge to the set

Input example:
4 5 1 2 1 1 3 2 1 4 3 2 3 2 3 4 4
Output example:
6
#include <iostream>
#include <cstring>
#include <algorithm>
using namespace std;
const int N = 100010, M = 200010, INF = 0x3f3f3f3f;
int n, m;
int p[N];
struct Edge{
int a, b, w;
bool operator< (const Edge &W)const
{
return w < W.w;
}
}edges[M];
int find(int x) //Joint search set (compressed path)
{
if (p[x] != x) p[x] = find(p[x]);
return p[x];
}
int kruskal()
{
sort(edges, edges + m);
for (int i = 1; i <= n; i ++ ) p[i] = i; // Initialize and query set
int res = 0, cnt = 0;
for (int i = 0; i < m; i ++ ){
int a = edges[i].a, b = edges[i].b, w = edges[i].w;
a = find(a), b = find(b);
if (a != b){
p[a] = b;
res += w;
cnt ++ ;
}
}
if (cnt < n-1) return INF;
else return res;
}
int main()
{
cin >> n >> m;
for (int i = 0; i < m; i ++ )
{
int a, b, w;
scanf("%d%d%d", &a, &b, &w);
edges[i] = {a, b, w};
}
int t = kruskal();
if (t == INF) cout << "impossible";
else cout << t;
return 0;
}
 https://www.acwing.com/problem/content/851/
https://www.acwing.com/problem/content/851/