Here we discuss six mistakes that novices are prone to make. These mistakes have nothing to do with the API or syntax of the tools you use, but are directly related to your level of knowledge and experience. In practice, if these problems occur, there may not be any error prompt, but it will bring us great trouble in application.
Read large files using the functions provided by pandas
The first error is related to actually using pandas to complete some tasks. Specifically, the data sets of tables we actually deal with are very large. Read using pandas_ CSV reading large files will be your biggest error.
Why? Because it's too slow! Looking at this test, we load the TPS October dataset, which has 1M rows and about 300 features, occupying 2.2GB of disk space.
import pandas as pd
%%time
tps_october = pd.read_csv("data/train.csv")
Wall time: 21.8 s
read_csv took about 22 seconds. You might say 22 seconds is not much. However, in a project, many experiments need to be performed at different stages. We will create many separate scripts for cleaning, feature engineering, model selection, and other tasks. Waiting for data to load for 20 seconds becomes very long. In addition, the dataset may be larger and take longer. So what is the faster solution?
The solution is to abandon Pandas at this stage and use other alternatives designed for fast IO. My favorite is datatable, but you can also choose Dask, Vaex, cuDF, etc. Here is the time required to load the same dataset with datatable:
import datatable as dt # pip install datatble
%%time
tps_dt_october = dt.fread("data/train.csv").to_pandas()
------------------------------------------------------------
Wall time: 2 s
Only 2 seconds, 10 times the gap
No Vectorization
One of the most important rules in functional programming is never to use loops. It seems that adhering to this "acyclic" rule when using Pandas is the best way to speed up the calculation.
Functional programming uses recursion instead of loops. Although there will be various problems with recursion (we don't consider this here), vectorization is the best choice for scientific computing!
Vectorization is the core of pandas and NumPy, which performs mathematical operations on the entire array rather than a single scalar. Pandas already has a wide range of vectorization functions. We don't need to reinvent the wheel, just focus on how to calculate.
Most of the arithmetic operators (+, -, *, /, * *) that perform Python in Pandas work in vectorization. In addition, any other mathematical functions seen in Pandas or NumPy have been vectorized.
To verify the speed improvement, we will use the following big_function, which takes three columns as input and performs some meaningless arithmetic as test:
def big_function(col1, col2, col3):
return np.log(col1 ** 10 / col2 ** 9 + np.sqrt(col3 ** 3))
First, we use this function with apply, the fastest iterator in Pandas:
%time tps_october['f1000'] = tps_october.apply(
lambda row: big_function(row['f0'], row['f1'], row['f2']), axis=1
)
-------------------------------------------------
Wall time: 20.1 s
The operation took 20 seconds. Let's use the core NumPy array in a vectorized way to do the same:
%time tps_october['f1001'] = big_function(tps_october['f0'].values,
tps_october['f1'].values,
tps_october['f2'].values)
------------------------------------------------------------------
Wall time: 82 ms
It took only 82 milliseconds, about 250 times faster.
In fact, we can't completely abandon the cycle. Because not all data operations are mathematical operations. But whenever you find that you need to use some loop functions (such as apply, applymap, or itertubles), it's a good habit to take a moment to see if what you want to do can be vectorized.
Data types, dtypes!
We can specify the data type according to the memory usage.
The worst and most memory consuming data type in Pandas is object, which also limits some functions of Pandas. We also have floating point numbers and integers for the rest. The following table shows all types of Pandas:
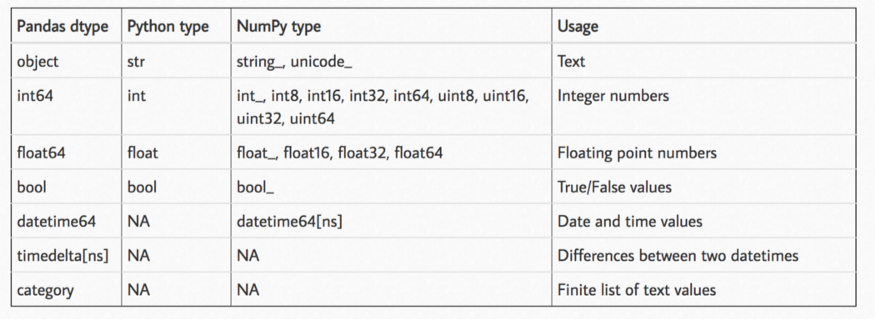
In Pandas naming scheme, the number after the data type name indicates how many bits of memory each number in this data type will occupy. Therefore, the idea is to convert each column in the dataset to a subtype as small as possible. We just need to judge according to the rules. This is the rule table:

Generally, floating point numbers are converted to float16/32 and columns with positive and negative integers are converted to int8/16/32 according to the above table. uint8 can also be used for Booleans and positive integers only to further reduce memory consumption.
You must be familiar with this function, because it is widely used in Kaggle. It converts floating-point numbers and integers to their minimum child types according to the above table:
def reduce_memory_usage(df, verbose=True):
numerics = ["int8", "int16", "int32", "int64", "float16", "float32", "float64"]
start_mem = df.memory_usage().sum() / 1024 ** 2
for col in df.columns:
col_type = df[col].dtypes
if col_type in numerics:
c_min = df[col].min()
c_max = df[col].max()
if str(col_type)[:3] == "int":
if c_min > np.iinfo(np.int8).min and c_max < np.iinfo(np.int8).max:
df[col] = df[col].astype(np.int8)
elif c_min > np.iinfo(np.int16).min and c_max < np.iinfo(np.int16).max:
df[col] = df[col].astype(np.int16)
elif c_min > np.iinfo(np.int32).min and c_max < np.iinfo(np.int32).max:
df[col] = df[col].astype(np.int32)
elif c_min > np.iinfo(np.int64).min and c_max < np.iinfo(np.int64).max:
df[col] = df[col].astype(np.int64)
else:
if (
c_min > np.finfo(np.float16).min
and c_max < np.finfo(np.float16).max
):
df[col] = df[col].astype(np.float16)
elif (
c_min > np.finfo(np.float32).min
and c_max < np.finfo(np.float32).max
):
df[col] = df[col].astype(np.float32)
else:
df[col] = df[col].astype(np.float64)
end_mem = df.memory_usage().sum() / 1024 ** 2
if verbose:
print(
"Mem. usage decreased to {:.2f} Mb ({:.1f}% reduction)".format(
end_mem, 100 * (start_mem - end_mem) / start_mem
)
)
return df
Let's use it on the TPS October data to see how much we can reduce:
>>> reduce_memory_usage(tps_october) Mem. usage decreased to 509.26 Mb (76.9% reduction)
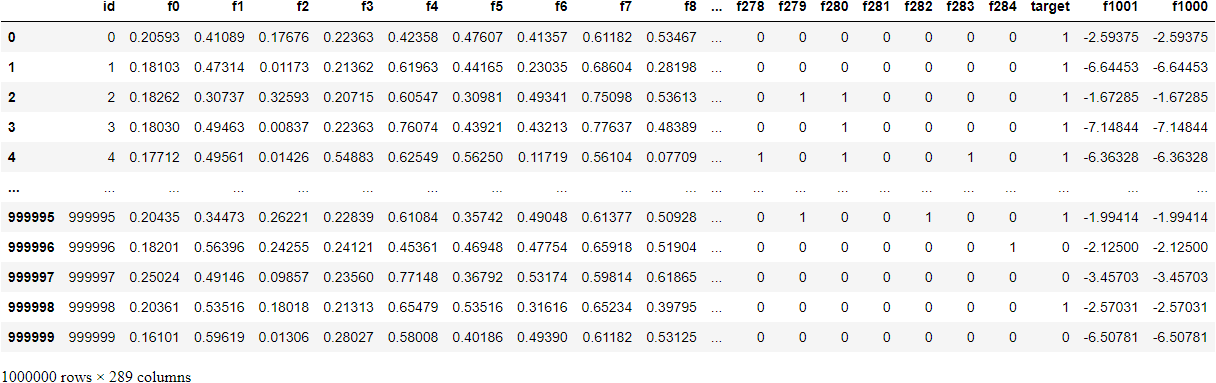
We compressed the data set from the original 2.2GB to 510MB. When we save df to csv file, this reduction in memory consumption will be lost, because csv is still saved in the form of string, but if we use pickle to save, it will be no problem.
Why reduce memory footprint? When using large machine learning model to process such data sets, the occupation and consumption of memory play an important role. Once you encounter some OutOfMemory errors, you will start to catch up and learn such skills to keep the computer working happily (who forced Kaggle to give only 16G of memory).
No style
One of the most wonderful features of Pandas is that it can set different styles when displaying DF, and render the original DF as an HTML table with some CSS in Jupiter.
Pandas allows its DataFrame to be styled through the style property.
tps_october.sample(20, axis=1).describe().T.style.bar(
subset=["mean"], color="#205ff2"
).background_gradient(subset=["std"], cmap="Reds").background_gradient(
subset=["50%"], cmap="coolwarm"
)
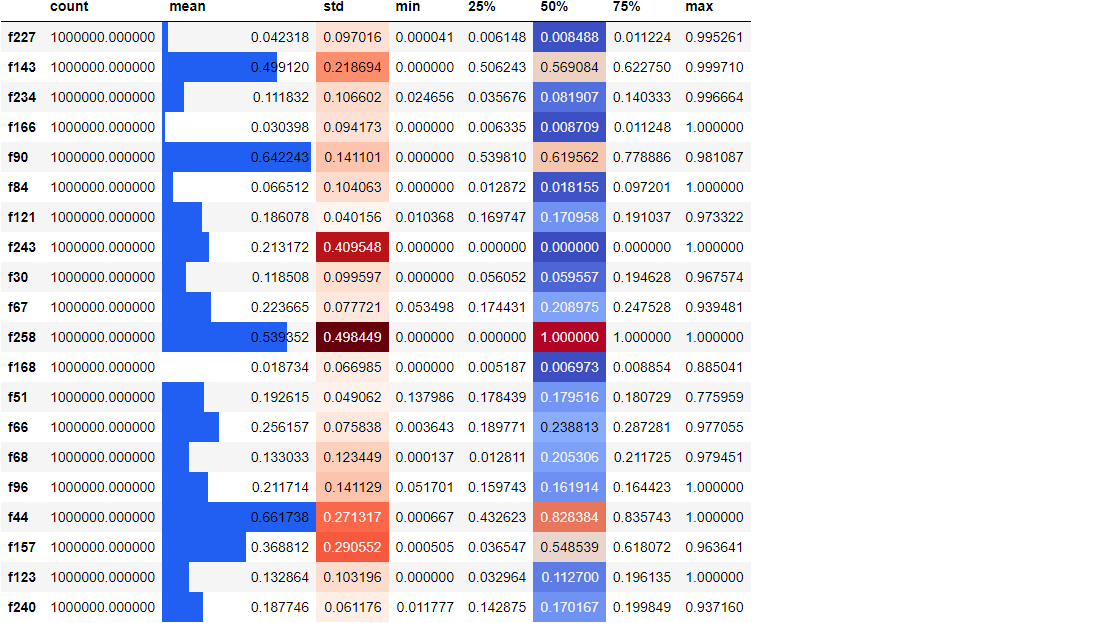
We randomly select 20 columns, create a 5-digit summary for them, transpose the results, and color the mean, standard deviation and median columns according to their size. Adding such a style can make it easier for us to find the pattern in the original number and set it without using other visualization libraries.
In fact, there is nothing wrong with not styling df. But it's really a good feature, isn't it.
Save the file in CSV format
Just as reading CSV files is very slow, so is saving data back to them. The following is the time required to save TPS October data to CSV:
%%time
tps_october.to_csv("data/copy.csv")
------------------------------------------
Wall time: 2min 43s
It took nearly three minutes. To save time, you can save it as parquet, feather or even pickle.
%%time
tps_october.to_feather("data/copy.feather")
Wall time: 1.05 s
--------------------------------------------------------------------------------
%%time
tps_october.to_parquet("data/copy.parquet")
Wall time: 7.84 s
Don't look at the document!
In fact, the most serious mistake for me was not reading Pandas's documents. But normally no one reads the document, right. Sometimes we would rather search on the Internet for hours than read documents.
But when it comes to Pandas, this is a very big mistake. Because it has an excellent user guide like sklearn, covering everything from basic knowledge to how to contribute code, and even how to set more beautiful themes (maybe because there are too many, so no one sees it).
All the errors I mentioned today can be found in the documentation. Even in the "large dataset" section of the document, you will be specifically told to use other software packages (such as Dask) to read large files and stay away from Pandas. In fact, if I had time to read the user guide from beginning to end, I might put forward 50 novice errors, so I'd better look at the documentation.
summary
Today, we learned the six most common mistakes novices make when using Pandas.
Most of the errors we mentioned here are related to large data sets, which may occur only when using GB data sets. If you're still dealing with novice data sets like Titanic, you may not feel these problems. But when you start working with real-world data sets, these concepts will make others feel that you are not a novice, but a person with real experience.
Author: Bex T